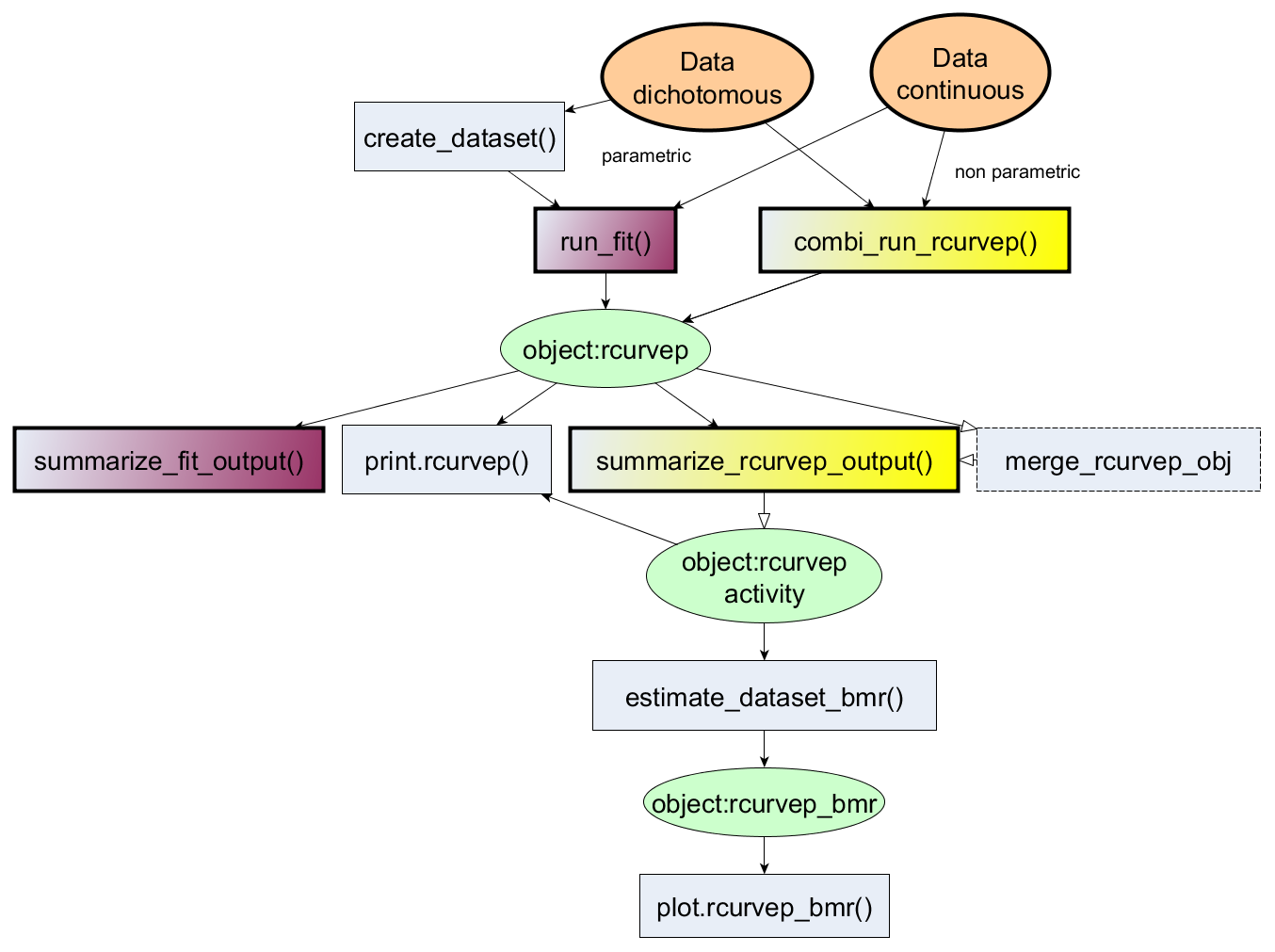
The package provides an R interface for processing concentration-response datasets using Curvep, a response noise filtering algorithm. The algorithm was described in the publications (Sedykh A et al. (2011) doi:10.1289/ehp.1002476 and Sedykh A (2016) doi:10.1007/978-1-4939-6346-1_14). Other parametric fitting approaches (e.g., Hill equation) are also adopted for ease of comparison. Also, methods for calculating the confidence interval around the activity metrics are also provided. The methods are based on the bootstrap approach to simulate the datasets (Hsieh J-H et al. doi:10.1093/toxsci/kfy258). The simulated datasets can be used to derive the baseline noise threshold in an assay endpoint. This threshold is critical in the toxicological studies to derive the point-of-departure (POD).
# the development version from GitHub:
# install.packages("devtools")
devtools::install_github("moggces/Rcurvep")
devtools::install_github("moggces/Rcurvep", build_vignettes = TRUE)
library(Rcurvep)
data("zfishbeh")
out_curvep <- combi_run_rcurvep(zfishbeh, TRSH = 30) # using Curvep with BMR = 30
out_fit <- run_fit(zfishbeh) data("zfishdev_act")
out_bmr <- estimate_dataset_bmr(zfishdev_act)## $`1`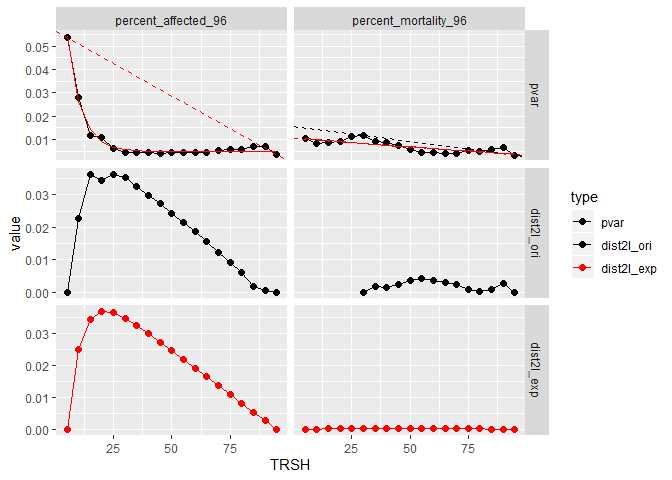
To learn more about Rcurvep, start with the vignettes:
browseVignettes(package = "Rcurvep")