IRTest can be a useful tool for IRT (item response
theory) parameter estimation, especially when the violation of normality
assumption on latent distribution is suspected.
IRTest deals with uni-dimensional latent
variable.
In IRTest, along with the conventional approach that
assumes normality on latent distribution, several methods can be applied
for estimation of latent distribution:
+ empirical histogram method,
+ two-component Gaussian mixture distribution,
+ Davidian curve,
+ kernel density estimation.
You can install IRTest on R-console with:
install.packages("IRTest")Followings are functions of IRTest available for users.
IRTest_Dich is the estimation function when all
items are dichotomously scored.
IRTest_Poly is the estimation function when all
items are polytomously scored.
IRTest_Mix is the estimation function for a
mixed-format test, a combination of dichotomous item(s) and polytomous
item(s).
DataGeneration generates several objects that are
useful for computer simulation studies. Among these are starting values
for an algorithm and artificial item-response data that can be passed to
IRTest_Dich, IRTest_Poly, or
IRTest_Mix
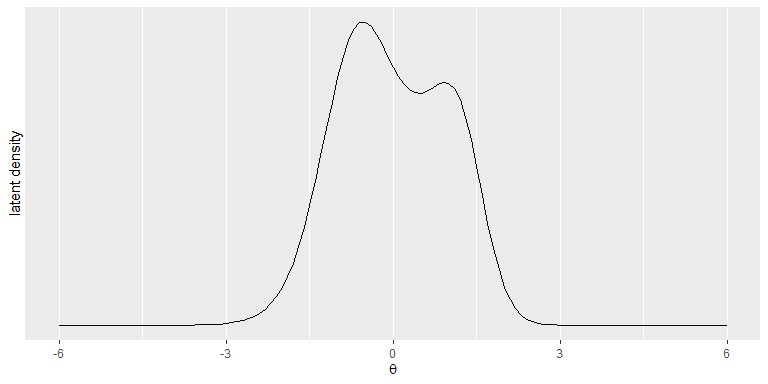
plot_LD draws a plot of the estimated latent
distribution.
dist2 is a probability density function of
two-component Gaussian mixture distribution.
original_par_2GM converts re-parameterized
parameters of two-component Gaussian mixture distribution into original
parameters.
A simulation study for a Rasch model can be done in following manners:
library(IRTest)Alldata <- DataGeneration(seed = 123456789,
model_D = rep(1, 10),
N=500,
nitem_D = 10,
nitem_P = 0,
d = 1.664,
sd_ratio = 2,
prob = 0.3)
data <- Alldata$data_D
item <- Alldata$item_D
initialitem <- Alldata$initialitem_D
theta <- Alldata$thetaFor an illustrative purpose, empirical histogram method is used for estimation of latent distribution.
Mod1 <- IRTest_Dich(initialitem = initialitem,
data = data,
model = rep(1, 10),
latent_dist = "EHM",
max_iter = 200,
threshold = .001)### True item parameters
item
#> [,1] [,2] [,3]
#> [1,] 1 -0.96 0
#> [2,] 1 0.67 0
#> [3,] 1 0.88 0
#> [4,] 1 0.55 0
#> [5,] 1 -0.20 0
#> [6,] 1 0.99 0
#> [7,] 1 0.38 0
#> [8,] 1 0.30 0
#> [9,] 1 1.93 0
#> [10,] 1 0.53 0
### Estimated item parameters
Mod1$par_est
#> a b c
#> [1,] 1 -0.7383615 0
#> [2,] 1 0.5071181 0
#> [3,] 1 0.7980726 0
#> [4,] 1 0.5274258 0
#> [5,] 1 -0.3909154 0
#> [6,] 1 0.8954094 0
#> [7,] 1 0.4264496 0
#> [8,] 1 0.3068586 0
#> [9,] 1 1.9525167 0
#> [10,] 1 0.4465375 0
### Plotting
par(mfrow=c(1,2))
plot(item[,2], Mod1$par_est[,2], xlab = "true", ylab = "estimated", main = "item parameters")
abline(a=0,b=1)
plot(theta, Mod1$theta, xlab = "true", ylab = "estimated", main = "ability parameters")
abline(a=0,b=1)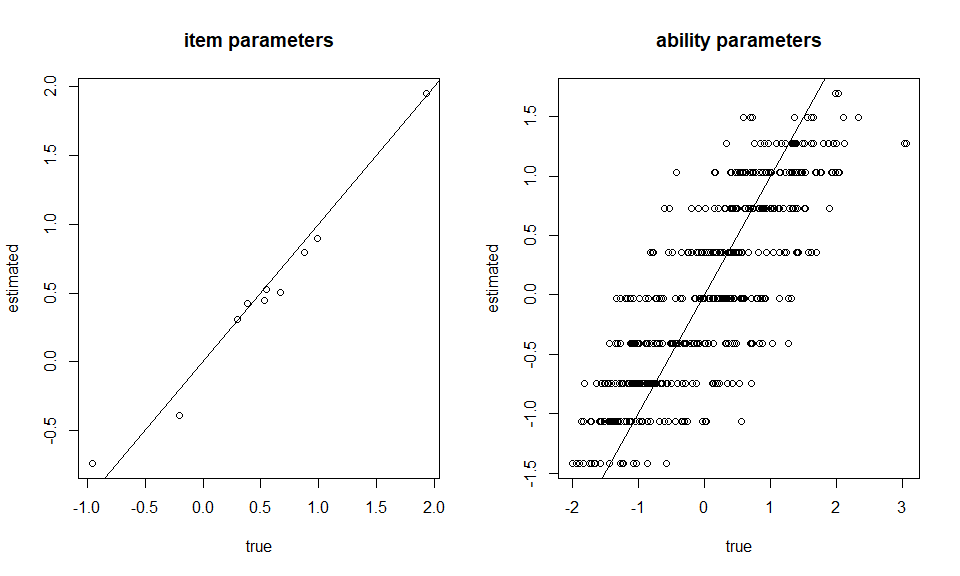
plot_LD(Mod1)